In a new article from our lab, Zisan Rahman tells us how to identify essential domains from Tn-seq data.
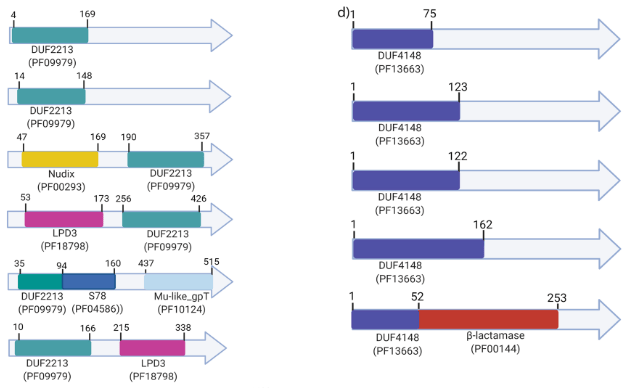
In bacteria, essential genes are usually identified by high‑density transposon mutagenesis followed by sequencing of insertion sites (Tn‑seq). These studies assign the term “essential” to whole genes rather than the protein domain sequences that encode the essential functions. However, genes can code for multiple protein domains that evolve their functions independently. Here, we developed an in silico pipeline to identify genes that encode essential domains. Together, our study suggests that essentiality should be assigned to individual protein domains rather than genes, contributing to a first functional characterization of protein domains of unknown function.